This tutorial is taken from the official Anvi’o infant gut metagenome tutorial.
We will try the “binning” mode of Anvi’o using a pre-made database based on the Sharon et al. 2013 infant gut metagenome, which contains 11 feacal samples from a premature infant collected between Day 15 and Day 24 of her life in a time-series design.
The dataset was co-assembled and processed the resulting contigs longer than 1K using anvi’o following the anvi’o metagenomic workflow.
Get the data
curl -L -o INFANT-GUT-TUTORIAL.tar.gz \
-H "User-Agent: Chrome/115.0.0.0" \
https://figshare.com/ndownloader/files/45076909
tar -zxvf INFANT-GUT-TUTORIAL.tar.gz && cd INFANT-GUT-TUTORIAL
This database was created using Anvi’o v6, so we need to upgrade the database files (.db) to be compatible with the latest Anvi’o version:
anvi-upgrade-database --migrate-safely *.db
Open the Anvi’o interface
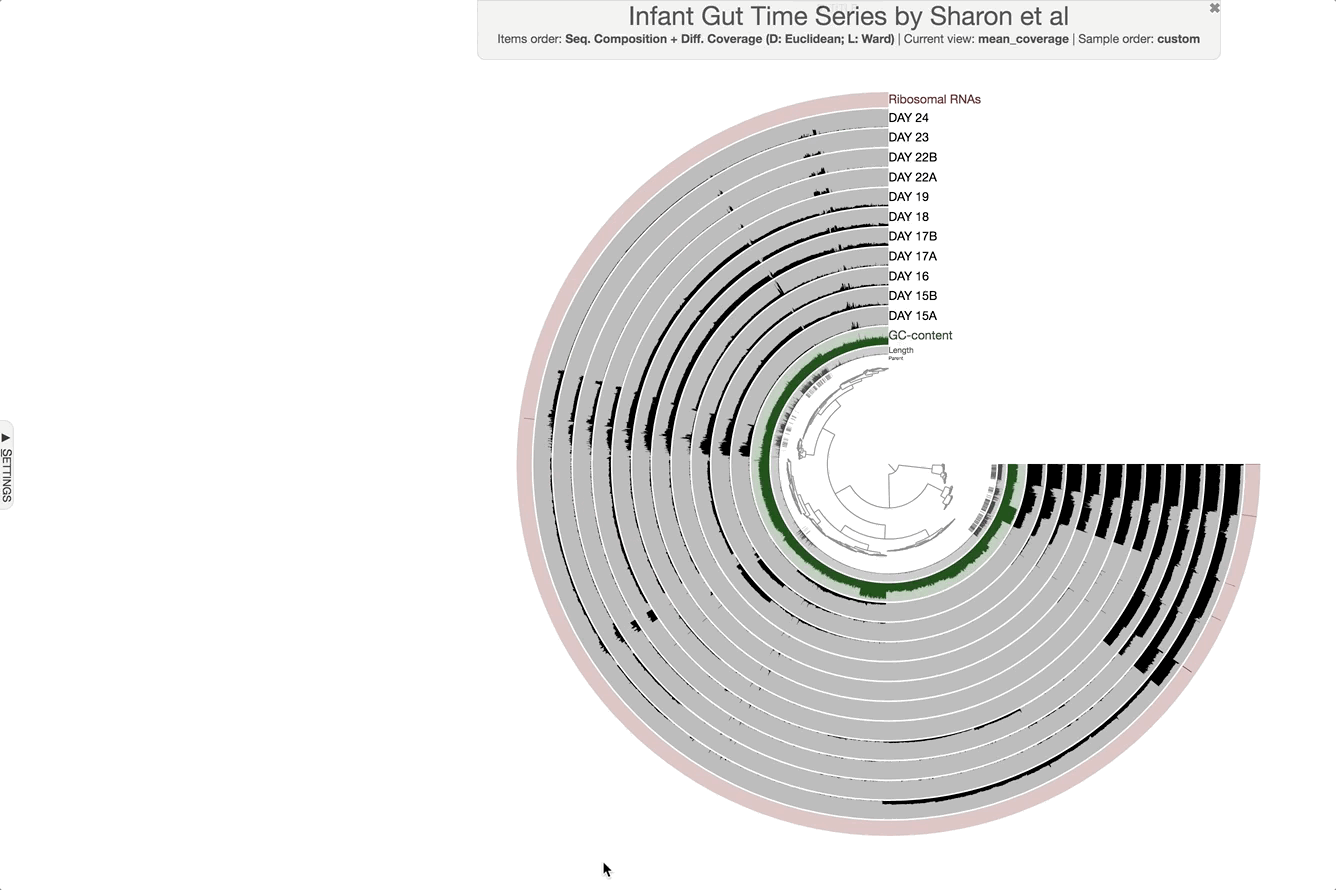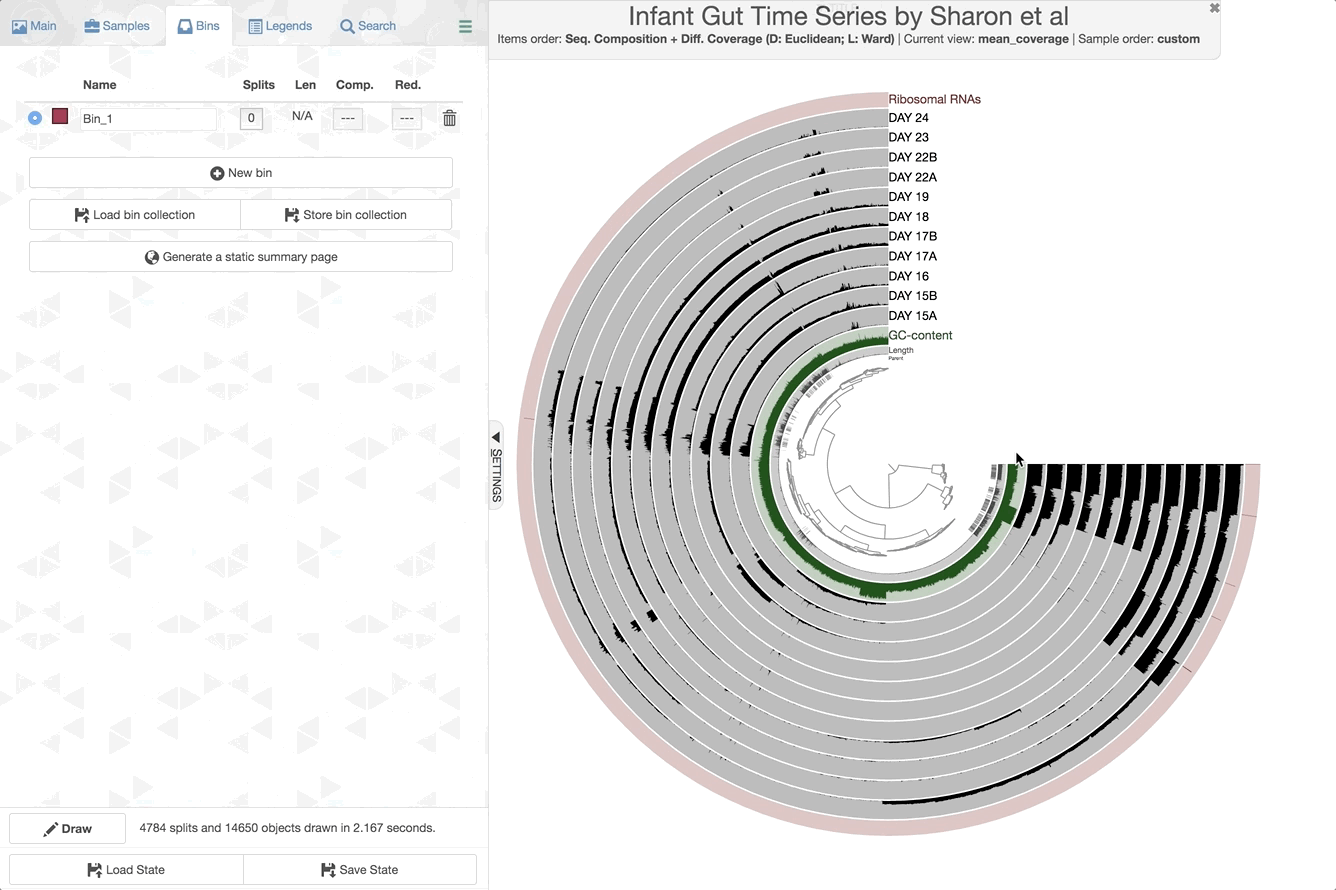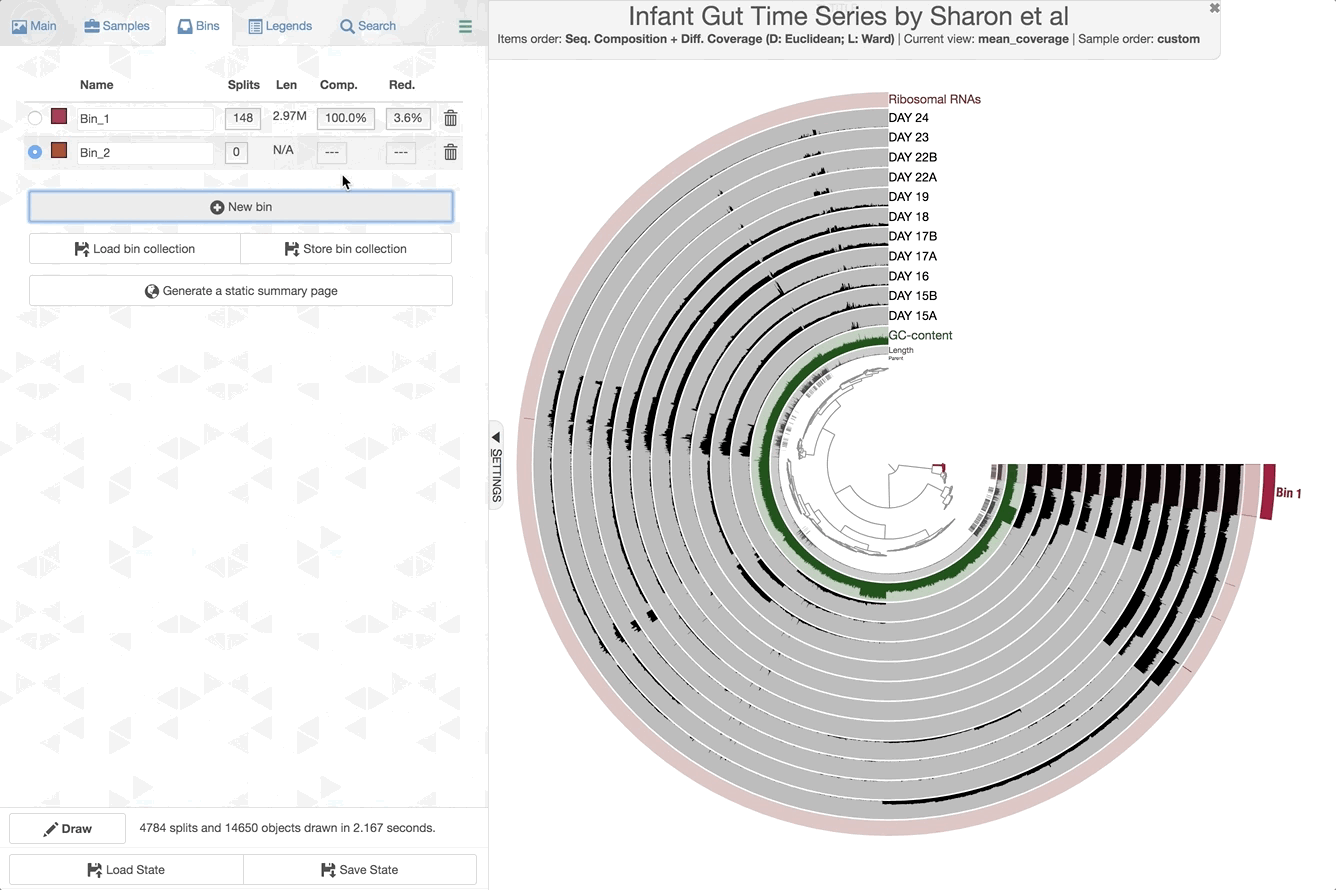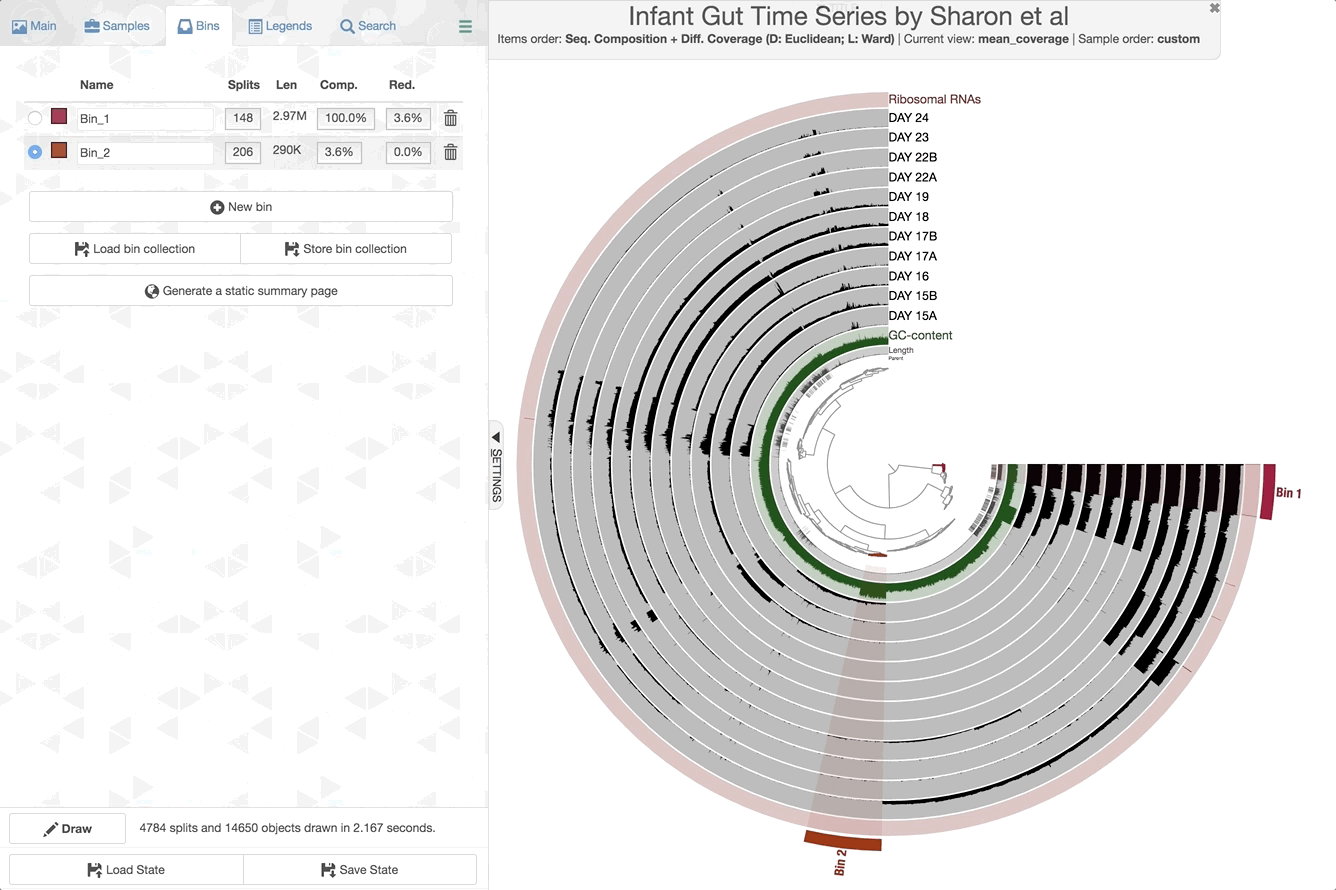
We can now check the data as produced:
anvi-interactive -p PROFILE.db -c CONTIGS.db
Estimating SCG taxonomy
Anvi’o has a database of single-copy core genes (SCGs) for taxonomic classification. We can use it to estimate the taxonomy of our bins:
anvi-run-scg-taxonomy -c CONTIGS.db --num-threads 4
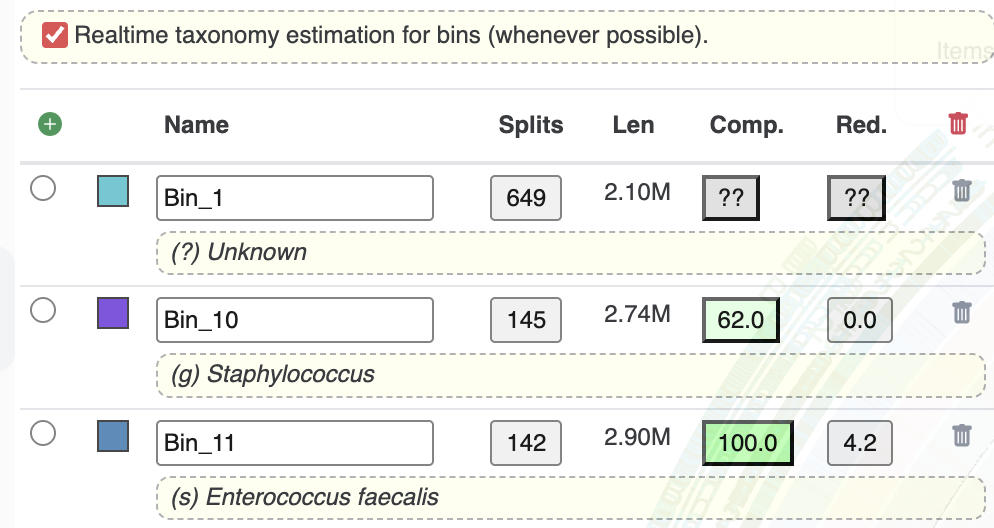
If now we re-open the interface, we can see the taxonomy estimates for each bin by clicking the “Realtime taxonomy” checkbox:
