This page is based on the official interface tutorial from Anvi’o.
Prepare a directory
Create a new directory and navigate into it to download a simple matrix dataset for practice:
mkdir anvio-interface
cd anvio-interface
# Download example dataset
wget http://merenlab.org/tutorials/interactive-interface/files/data.txt
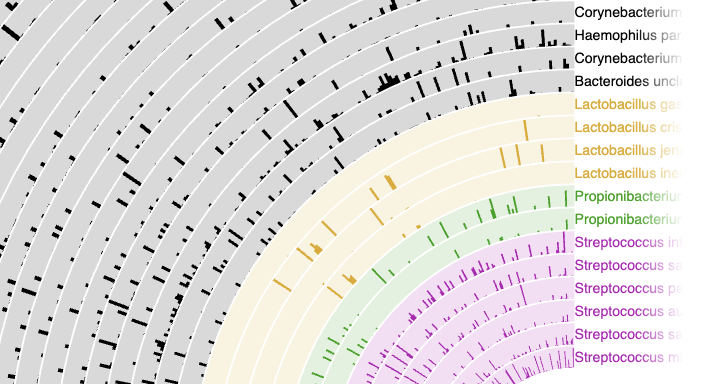
We can have a first look at the data:
less -S -x 20 data.txt
# or a quick preview (first 10 lines, first 6 columns)
head -n 10 data.txt | cut -f 1-6
Launch the interface
Activate the anvio-dev conda environment:
conda activate anvio-dev
then launch the Anvi’o interactive interface with:
anvi-interactive --manual -d data.txt \
-p profile.db \
--title "Taxonomic profiles of 690 gut metagenomes"
![]() What is “profile.db”?
What is “profile.db”?
![]() Using
Using anvi-interactive --help, can you find what --manual does?
This will open a new window in your web browser with the Anvi’o interactive interface.
You will see the panel with the controls, but to render the actual data you need to click on the “Render” button (green button on the top left).

-
 try to zoom and pan the view using your mouse or the controls on the left side of the interface.
try to zoom and pan the view using your mouse or the controls on the left side of the interface. -
🎨 try changing the colours of some layers
-
↖ try using the “Data” pane to inspect what the mouse is pointing at
Where is the dendrogram?
Our matrix does not have a dendrogram, because we did not provide any hierarchical clustering information.
However, we can generate a dendrogram based on the data in the matrix. This can be done using the anvi-matrix-to-newick command, that will produce a tree in Newick format.
anvi-matrix-to-newick data.txt \
-o tree.txt
# Wanna peek inside the output?
less tree.txt
Relaunch the interface with the tree
To use the tree we just generated, we need to relaunch the interface with the -t option:
anvi-interactive --manual -d data.txt \
-p profile.db\
--title "Taxonomic profiles of 690 gut metagenomes" \
--tree tree.txt
![]() So far we used a text file as input, in typical workflows we will generate special databases from genomics file.
So far we used a text file as input, in typical workflows we will generate special databases from genomics file.
Adding metadata
We can also add metadata to our dataset, for example to group samples by some characteristics.
For example:
| Metagenome | Body_Site | Body_Subsite | Host_Gender |
|---|---|---|---|
| SRS011061 | GastrointestinalTract | Stool | Female |
| SRS011090 | Oral | Buccal_mucosa | Female |
| SRS011098 | Oral | Supragingival_plaque | Female |
| SRS011126 | Oral | Supragingival_plaque | Male |
We can download a metadata file with this information:
wget http://merenlab.org/tutorials/interactive-interface/files/additional-items-data.txt
First, we need to import the metadata into Anvi’o:
anvi-import-misc-data additional-items-data.txt \
--target-data-table items \
--pan-or-profile-db profile.db
then we can re-launch the interface:
anvi-interactive -d data.txt \
-p profile.db \
--title "Taxonomic profiles of 690 HMP metagenomes" \
--tree tree.txt \
--manual
What’s next? There is an interesting tutorial on Trichodesmium, that will guide you through a complete workflow.