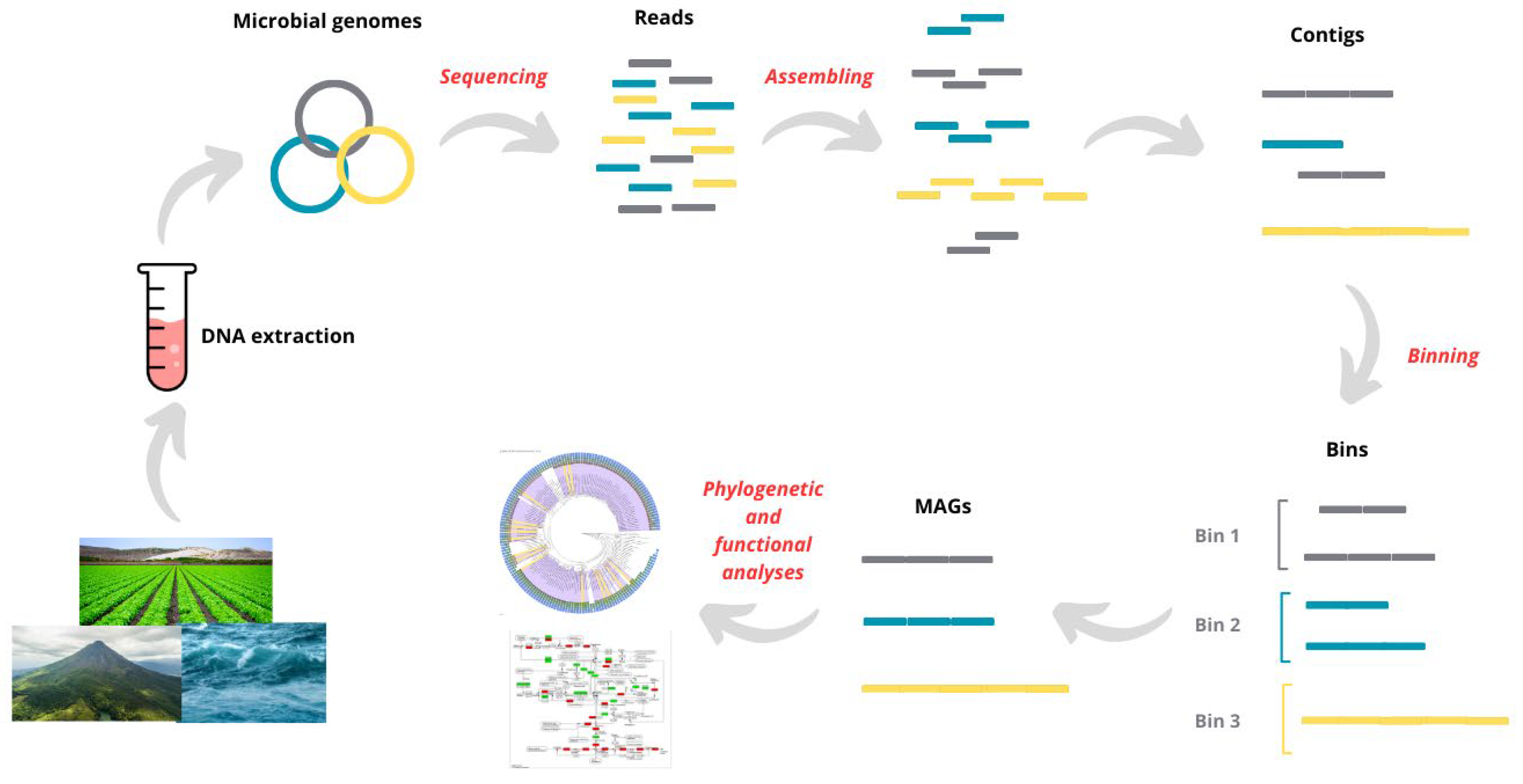
from Mirete, S (2025)
In this session we will see how to perform a de novo assembly of metagenomic reads using MEGAHIT (or Minia) and how to map back the reads to the assembled contigs using any short read mapper (e.g., Minimap2 or Bwa).
- ❓ What is the main difference between the assembly of isolate genomes and metagenomes?
- ❓ How can we evaluate the quality of a metagenomic assembly?
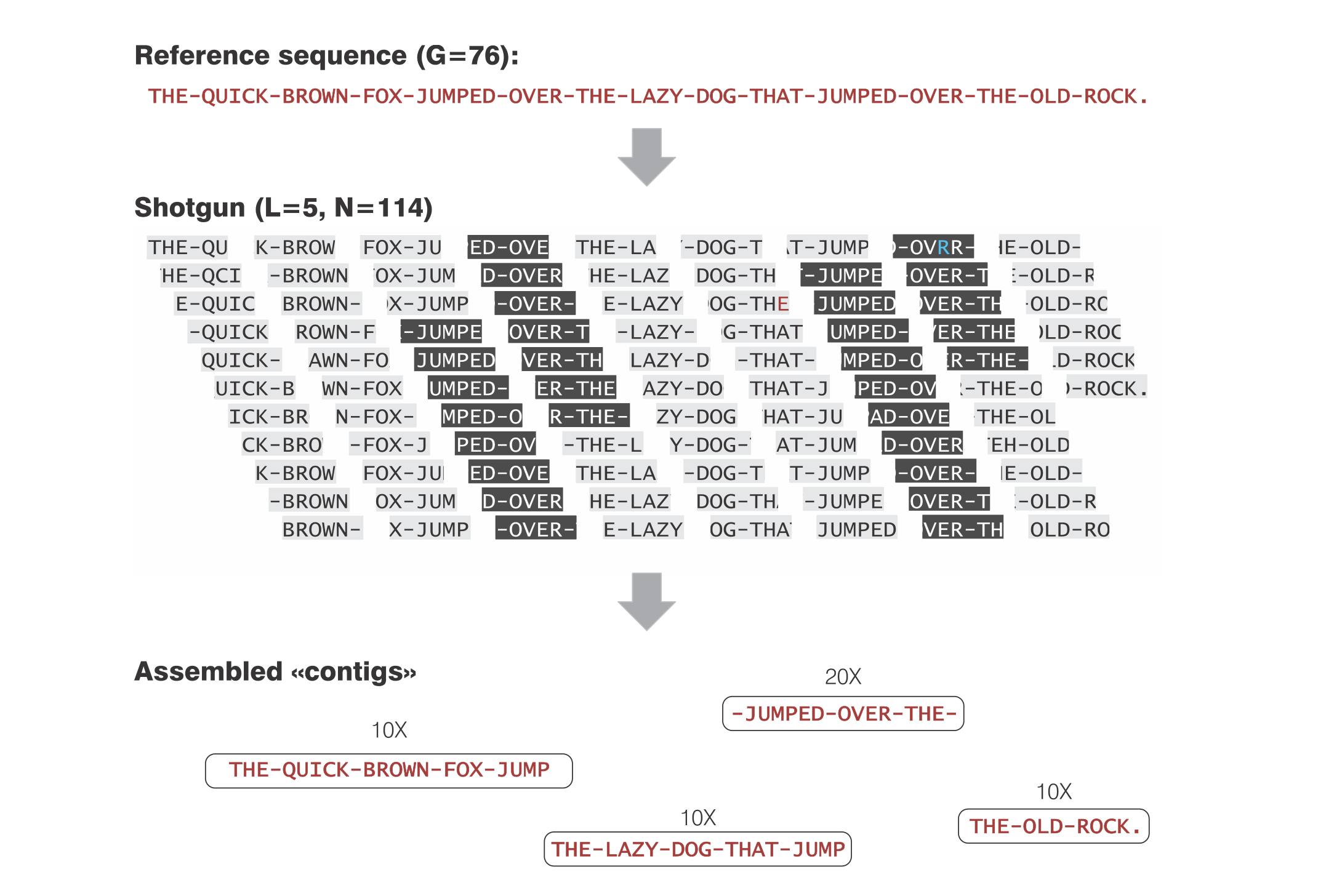
from Telatin, A (2014)
There are different approaches to assemble a metagenome. When ssing Illumina (short) reads, the only feasible way is to use de Bruijn graphs assemblers, which are based on the decomposition of reads into smaller k-mers.
- ❓ What are the advantages and disadvantages of using long reads for metagenomic assembly?
Next submodule:
Assembly tools for metagenomics